 
These birds need vegetables, as well as citrus peel, egg yolks, unprocessed grains, and turmeric to detoxify.Step 6) Analyzing your Mitochondrial DNA Sequences Trimming your mitochondrial DNA sequence using 4Peaks. In order to determine how related you are to your classmates, to Rarely Reclusive, and to the human ancient and modern groups in the Dolan DNA database, you will need to first determine whether or not your sequence is of sufficient quality for the analysis, and then to trim your sequence so that only good quality data is used. The 4Peaks program that you used during your yeast experiments will work very well for this purpose. Since 4Peaks is a Mac-only program, you will need to do the first two stages of this analysis on a Mac (your own, if you have one, or a departmental laptop if you do not). Examining the primary data of the mitochondrial control region You can then do the MEGA5 and Dolan DNA analysis on any computer platform. Open the “Mitochondrial DNA Sequences” folder and download your sequence file (labeled “TT(#)” to the desktop. Open the file using the program “4Peaks”. At this point you should see peaks of four different colors. There is also an X-axis that consists of a time unit of measurement, a nucleotide number, and a nucleotide (designated A, C, G, or T). 
There are a few things to note: o at many points there is only one peak and that the color of the peak determines the nucleotide indicated on the X-axis. The sequence at these points is of high quality, and can be used for phylogenetic analysis. O at some points, there is more than one peak, or the peak will look very broad or misshapen (see bases near the beginning, for example). This is due to technical problems and makes it difficult or impossible to predict the base that is at that position on the DNA strand. This is why there is occasionally a “N” instead of an A, C, G, or T. “N” designates that the identity of the base at this position was not clearly determined. Sequence with too many “Ns” cannot be used in phylogenetic analysis. 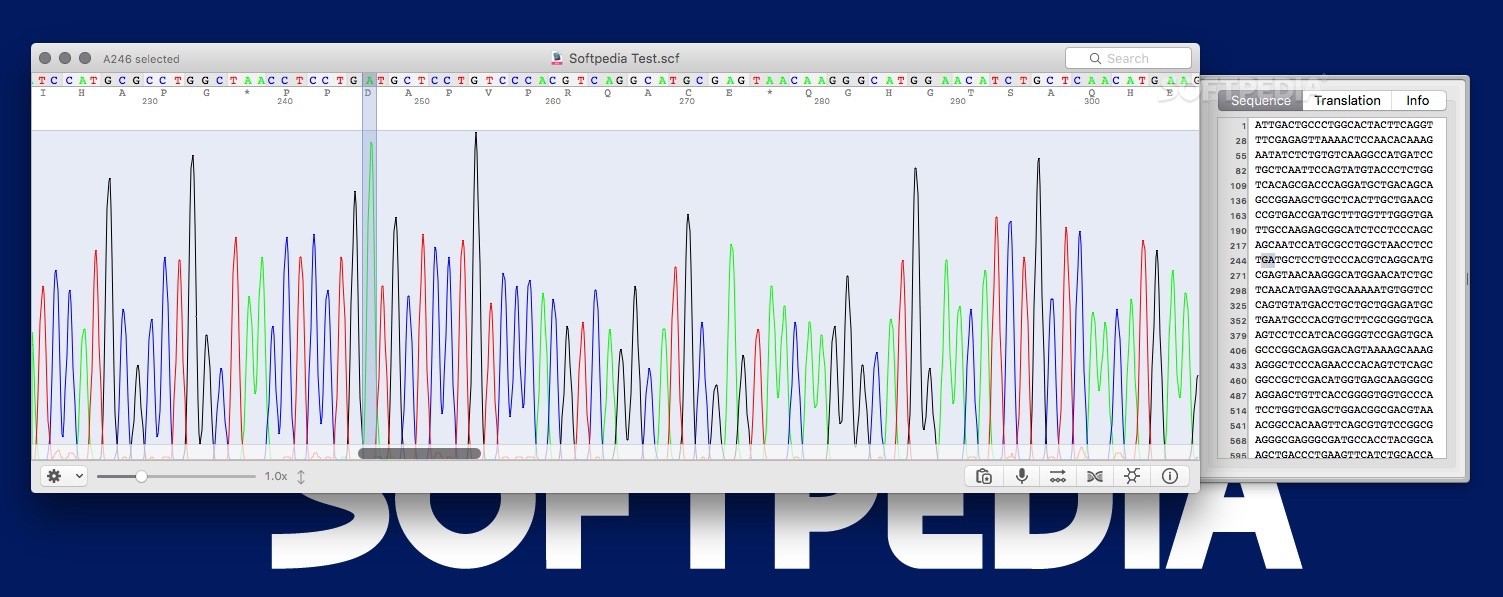
An example is shown below, although some sequences will certainly look even worse than this:Ĥ Peaks may also try to call sequences when the peaks are obviously of poor quality, as shown below. Areas with poor quality peaks should NOT be included in your sequence analysis. Does it have a section with good sequence, or are there N’s or poor quality peaks all the way through it? If there is good sequence available, highlight (select) the good sequence by clicking on the first base of the good stretch, and then holding and dragging to the right until you have selected the last base of the good stretch. If your sequence is not of sufficient quality, do not worry. Simply work with a sequence from your team that is. The sequence that remains in the 4Peaks window is now just the good quality DNA.ĭo NOT close your cropped file when you move to the next step – you will need it again soon! Select "Crop" from the drop-down menu in the lower left hand corner of the window (the ‘gear’ icon). Use “Export” to rename this cropped file and to save it to the desktop. fas This step must be done or the MEGA 6 program will not recognize the sequences. Repeat this process (cropping and exporting) for each member of your team. BLAST Analysis of your cropped sequence If a team member did not get good sequence of their own, simply choose one of the other class sequences.ĪT THIS STAGE, IF YOU ARE NOT YET USING YOUR OWN COMPUTER, EMAIL THE CROPPED FILES TO THE TEAM MEMBER WHO BROUGHT THE COMPUTER, AND THEN CONTINUE ON WITH THE 4PEAKS ANALYSIS BELOW.Ģ. Under the gear icon in the lower left corner. choose “BLAST Sequence” then Nucleotide (BLASTn). 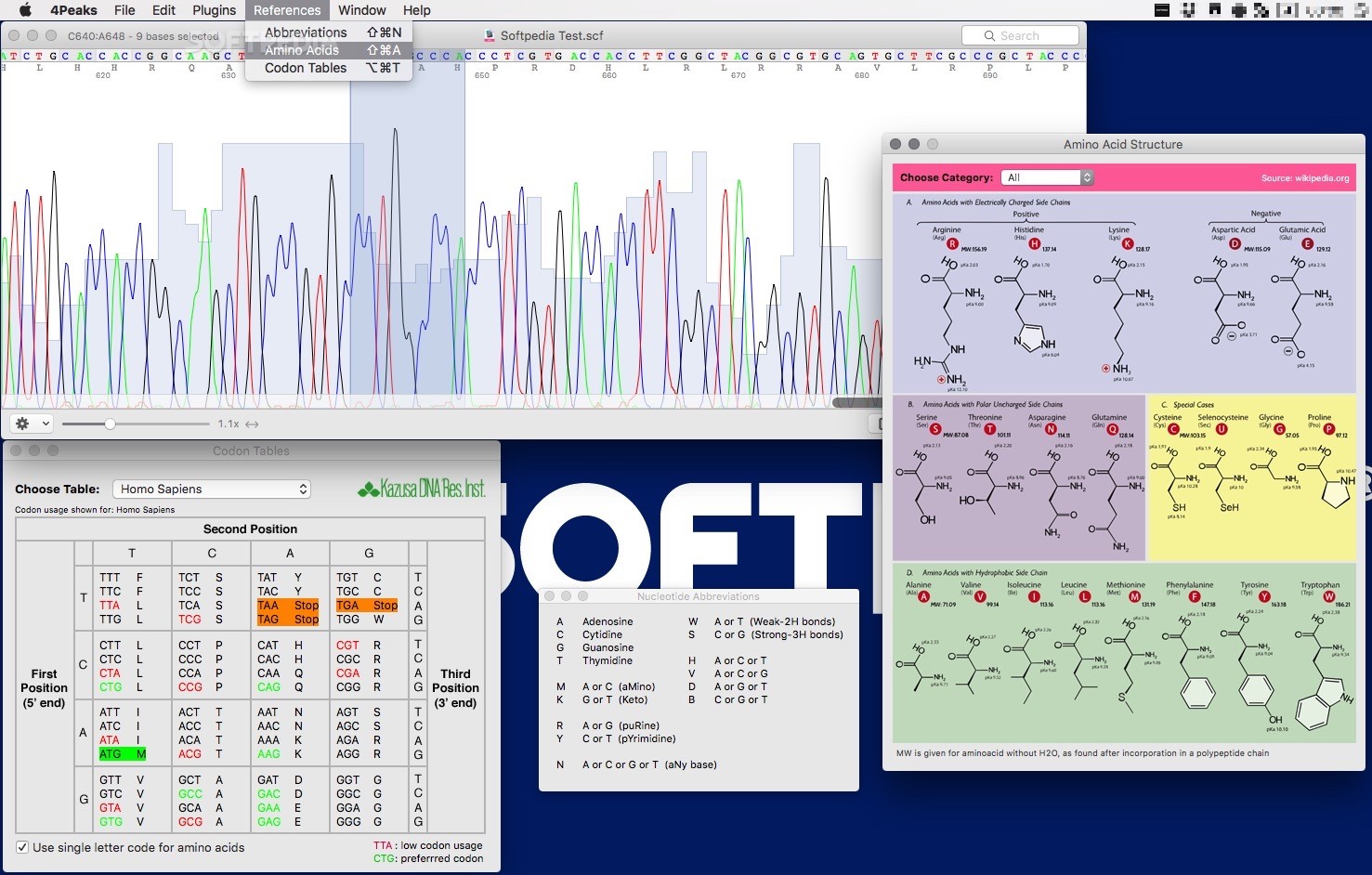
This is the same program that you used for your yeast DNA sequence comparisons, except we are now using the entire sequence database, rather than just yeast. You should see your sequence in the query window. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
